Synthetic control with pymc models#
import arviz as az
import causalpy as cp
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
seed = 42
Load data#
df = cp.load_data("sc")
treatment_time = 70
Donor pool selection#
Before fitting, it is good practice to inspect pairwise correlations among units in the pre-treatment period. Negatively correlated donors should be excluded to avoid interpolation bias [Abadie, 2021, Abadie et al., 2010].
pre = df.loc[:treatment_time]
corr, ax = cp.plot_correlations(pre, columns=["a", "b", "c", "d", "e", "f", "g"])
ax.set(title="Pre-treatment correlations")
[Text(0.5, 1.0, 'Pre-treatment correlations')]
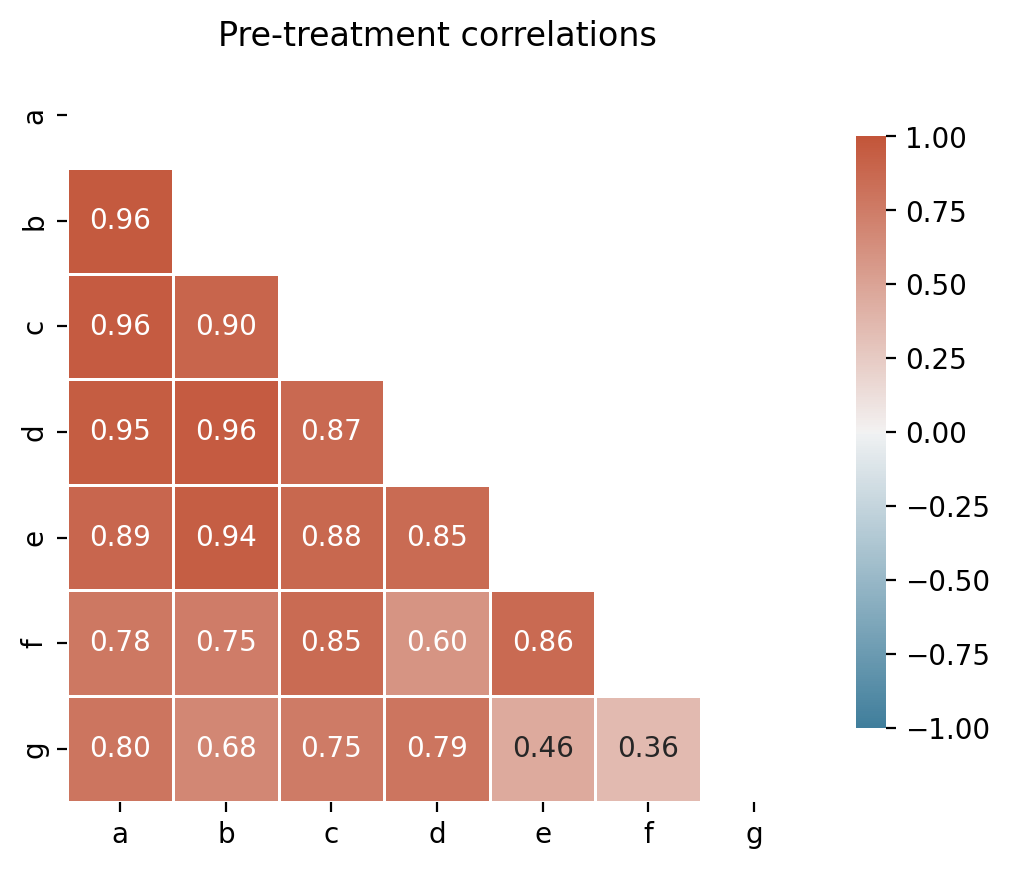
All control units are positively correlated with the treated unit (actual), so no donor pool curation is needed for this dataset. In practice, you would exclude any controls with negative or near-zero correlations.
Run the analysis#
Note
The random_seed keyword argument for the PyMC sampler is not necessary. We use it here so that the results are reproducible.
Convex Hull Assumption#
The synthetic control method uses non-negative weights that sum to one. This means the synthetic control is a convex combination of the control units—it can only produce values within the range spanned by the controls at each time point.
Note
For the method to work well, the treated unit’s pre-intervention trajectory should lie within the “envelope” of the control units. If all controls are consistently above or below the treated unit, the method cannot construct an accurate counterfactual.
CausalPy automatically checks this assumption when you create a SyntheticControl object and will issue a warning if violated. See the Convex hull condition glossary entry and Abadie et al. [2010] for more details. The Augmented Synthetic Control Method [Ben-Michael et al., 2021] can handle cases where this assumption is violated.
df.head()
| a | b | c | d | e | f | g | counterfactual | causal effect | actual | |
|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 0.806091 | -0.271100 | 0.664254 | 1.705922 | -1.449230 | 0.785439 | -1.234083 | -0.232966 | -0.0 | -0.076404 |
| 1 | 1.442677 | 0.462594 | 0.559940 | 1.862041 | -1.615366 | 0.386238 | -1.798470 | -0.069413 | -0.0 | -0.063946 |
| 2 | 1.916486 | 0.845719 | 0.894046 | 2.061073 | -1.978696 | 0.971803 | -0.490287 | 0.095440 | -0.0 | 0.392813 |
| 3 | 2.001511 | 1.067162 | 0.288606 | 1.934534 | -2.093951 | 1.889334 | -0.478545 | 0.261716 | -0.0 | 0.133106 |
| 4 | 2.662849 | 1.243252 | 0.966345 | 2.482698 | -1.407598 | 2.184504 | -0.434592 | 0.429432 | -0.0 | 0.293294 |
result = cp.SyntheticControl(
df,
treatment_time,
control_units=["a", "b", "c", "d", "e", "f", "g"],
treated_units=["actual"],
model=cp.pymc_models.WeightedSumFitter(
sample_kwargs={"target_accept": 0.95, "random_seed": seed}
),
)
Show code cell output
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [beta, y_hat_sigma]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 14 seconds.
Sampling: [beta, y_hat, y_hat_sigma]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
fig, ax = result.plot(plot_predictors=True)
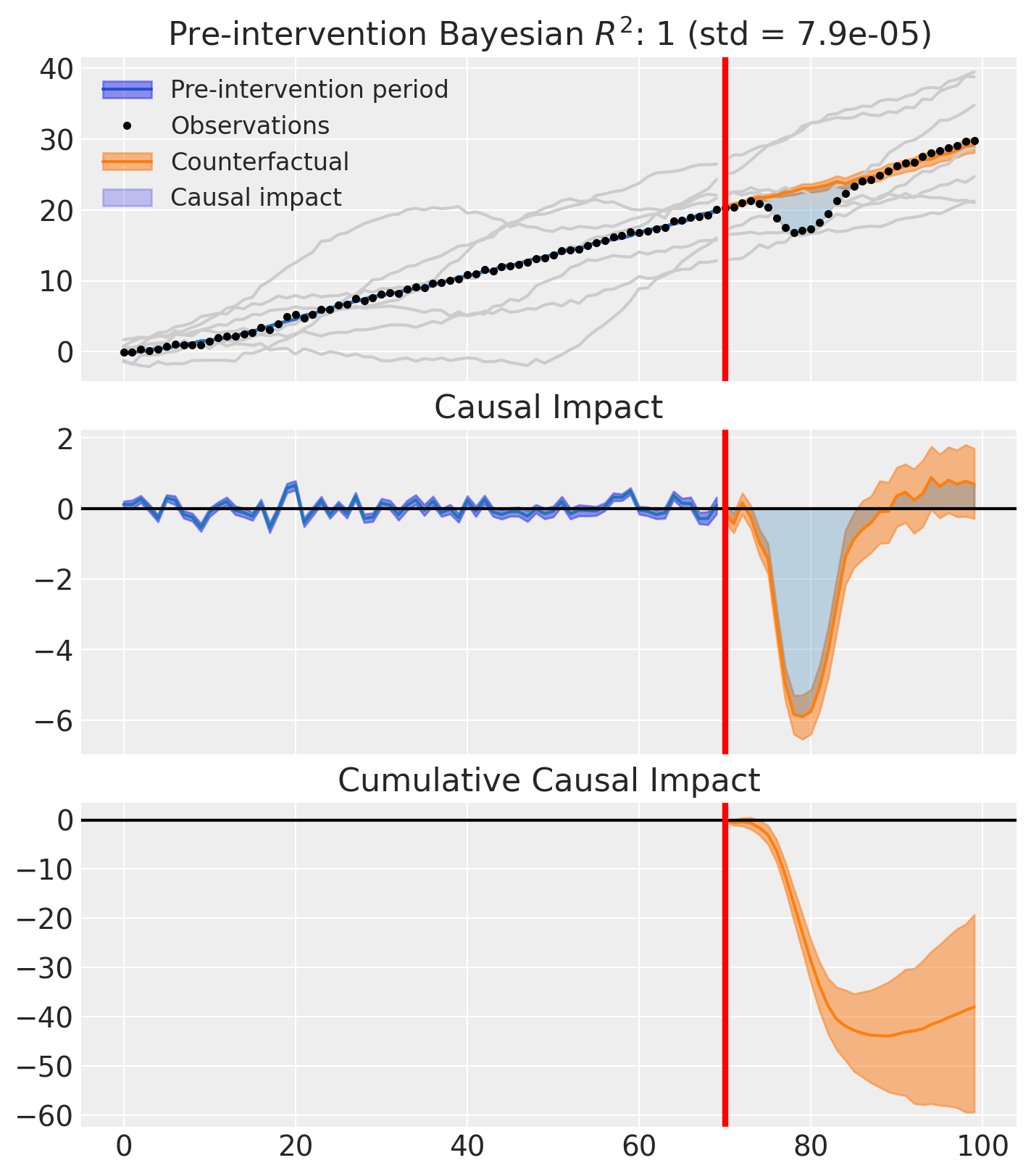
result.summary()
================================SyntheticControl================================
Control units: ['a', 'b', 'c', 'd', 'e', 'f', 'g']
Treated unit: actual
Model coefficients:
a 0.056, 94% HDI [0.0031, 0.14]
b 0.2, 94% HDI [0.14, 0.25]
c 0.21, 94% HDI [0.13, 0.29]
d 0.058, 94% HDI [0.0061, 0.12]
e 0.26, 94% HDI [0.23, 0.29]
f 0.11, 94% HDI [0.068, 0.15]
g 0.11, 94% HDI [0.074, 0.15]
y_hat_sigma 0.25, 94% HDI [0.21, 0.3]
Pre-treatment correlation (actual): 0.9992
We can get nicely formatted tables from our integration with the maketables package.
from maketables import ETable
result.set_maketables_options(hdi_prob=0.95)
ETable(result, coef_fmt="b:.3f \n [ci95l:.3f, ci95u:.3f]")
| y | |
|---|---|
| (1) | |
| coef | |
| a | 0.056 [0.000, 0.126] |
| b | 0.195 [0.142, 0.252] |
| c | 0.213 [0.130, 0.292] |
| d | 0.058 [0.001, 0.116] |
| e | 0.256 [0.224, 0.290] |
| f | 0.110 [0.067, 0.152] |
| g | 0.112 [0.073, 0.151] |
| stats | |
| N | 100 |
| Bayesian R2 | 0.998 |
| Format of coefficient cell: Coefficient [95% CI Lower, 95% CI Upper] | |
As well as the model coefficients, we might be interested in the average causal impact and average cumulative causal impact.
az.summary(result.post_impact.mean("obs_ind"))
| mean | sd | hdi_3% | hdi_97% | mcse_mean | mcse_sd | ess_bulk | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|---|
| x[actual] | -1.267 | 0.367 | -1.979 | -0.641 | 0.014 | 0.007 | 731.0 | 1495.0 | 1.01 |
Warning
Care must be taken with the mean impact statistic. It only makes sense to use this statistic if it looks like the intervention had a lasting (and roughly constant) effect on the outcome variable. If the effect is transient, then clearly there will be a lot of post-intervention period where the impact of the intervention has ‘worn off’. If so, then it will be hard to interpret the mean impacts real meaning.
We can also ask for the summary statistics of the cumulative causal impact.
# get index of the final time point
index = result.post_impact_cumulative.obs_ind.max()
# grab the posterior distribution of the cumulative impact at this final time point
last_cumulative_estimate = result.post_impact_cumulative.sel({"obs_ind": index})
# get summary stats
az.summary(last_cumulative_estimate)
| mean | sd | hdi_3% | hdi_97% | mcse_mean | mcse_sd | ess_bulk | ess_tail | r_hat | |
|---|---|---|---|---|---|---|---|---|---|
| x[actual] | -38.005 | 11.019 | -59.384 | -19.237 | 0.407 | 0.211 | 731.0 | 1495.0 | 1.01 |
Effect Summary Reporting#
For decision-making, you often need a concise summary of the causal effect with key statistics. The effect_summary() method provides a decision-ready report with average and cumulative effects, HDI intervals, tail probabilities, and relative effects.
# Generate effect summary for the full post-period
stats = result.effect_summary(treated_unit="actual")
stats.table
| mean | median | hdi_lower | hdi_upper | p_gt_0 | relative_mean | relative_hdi_lower | relative_hdi_upper | |
|---|---|---|---|---|---|---|---|---|
| average | -1.266830 | -1.269718 | -2.001737 | -0.613731 | 0.00075 | -5.155195 | -7.941135 | -2.576623 |
| cumulative | -38.004895 | -38.091551 | -60.052099 | -18.411916 | 0.00075 | -5.155195 | -7.941135 | -2.576623 |
print(stats.text)
During the Post-period (70 to 99), the response variable had an average value of approx. 23.21. By contrast, in the absence of an intervention, we would have expected an average response of 24.47. The 95% interval of this counterfactual prediction is [23.82, 25.21]. Subtracting this prediction from the observed response yields an estimate of the causal effect the intervention had on the response variable. This effect is -1.27 with a 95% interval of [-2.00, -0.61].
Summing up the individual data points during the Post-period, the response variable had an overall value of 696.16. By contrast, had the intervention not taken place, we would have expected a sum of 734.17. The 95% interval of this prediction is [714.58, 756.22].
The 95% HDI of the effect [-2.00, -0.61] does not include zero. The posterior probability of a decrease is 0.999. Relative to the counterfactual, the effect represents a -5.16% change (95% HDI [-7.94%, -2.58%]).
This analysis assumes that the control units used to construct the synthetic counterfactual were not themselves affected by the intervention, and that the pre-treatment relationship between control and treated units remains stable throughout the post-treatment period. We recommend inspecting model fit, examining pre-intervention trends, and conducting sensitivity analyses (e.g., placebo tests) to support any causal conclusions drawn from this analysis.
You can customize the summary in several ways:
Window: Analyze a specific time period instead of the full post-period
Direction: Specify whether you’re testing for an increase, decrease, or two-sided effect
Options: Include/exclude cumulative or relative effects
# Example: Analyze first half of post-period
post_indices = result.datapost.index
window_start = post_indices[0]
window_end = post_indices[len(post_indices) // 2]
stats_windowed = result.effect_summary(
window=(window_start, window_end),
treated_unit="actual",
direction="two-sided",
cumulative=True,
relative=True,
)
stats_windowed.table
| mean | median | hdi_lower | hdi_upper | p_two_sided | prob_of_effect | relative_mean | relative_hdi_lower | relative_hdi_upper | |
|---|---|---|---|---|---|---|---|---|---|
| average | -2.672167 | -2.655041 | -3.192889 | -2.168353 | 0.0 | 1.0 | -11.888489 | -13.89799 | -9.878958 |
| cumulative | -42.754673 | -42.480652 | -51.086217 | -34.693647 | 0.0 | 1.0 | -11.888489 | -13.89799 | -9.878958 |
print(stats_windowed.text)
During the Post-period (70 to 85), the response variable had an average value of approx. 19.78. By contrast, in the absence of an intervention, we would have expected an average response of 22.45. The 95% interval of this counterfactual prediction is [21.95, 22.97]. Subtracting this prediction from the observed response yields an estimate of the causal effect the intervention had on the response variable. This effect is -2.67 with a 95% interval of [-3.19, -2.17].
Summing up the individual data points during the Post-period, the response variable had an overall value of 316.49. By contrast, had the intervention not taken place, we would have expected a sum of 359.25. The 95% interval of this prediction is [351.19, 367.58].
The 95% HDI of the effect [-3.19, -2.17] does not include zero. The posterior probability of an effect is 1.000. Relative to the counterfactual, the effect represents a -11.89% change (95% HDI [-13.90%, -9.88%]).
This analysis assumes that the control units used to construct the synthetic counterfactual were not themselves affected by the intervention, and that the pre-treatment relationship between control and treated units remains stable throughout the post-treatment period. We recommend inspecting model fit, examining pre-intervention trends, and conducting sensitivity analyses (e.g., placebo tests) to support any causal conclusions drawn from this analysis.
Sensitivity analysis#
This section walks through placebo-in-time diagnostics for synthetic control using the pipeline API (EstimateEffect → SensitivityAnalysis → GenerateReport). For the shared workflow concepts — Pipeline, pluggable checks, and HTML reports — see Pipeline Workflow. For the full menu of checks and how they relate, see Sensitivity Checks in Pipeline Workflows.
Placebo-in-time check#
PlaceboInTime is the synthetic control default check registered in cp.SensitivityAnalysis.default_for(cp.SyntheticControl). It answers a falsification question central to synthetic control practice: if we pretended the intervention happened earlier, would we still see an effect? Placebo and falsification exercises are a standard component of synthetic-control credibility [Abadie, 2021].
How it works. The check shifts the treatment time backward into the pre-intervention period, fitting n_folds placebo experiments. From each fold it extracts the posterior cumulative impact, then fits a hierarchical Bayesian status-quo model that pools across folds to characterise the distribution of effects when no real intervention occurred. The actual cumulative impact is compared against this learned null, and the check passes when the posterior probability that the real effect lies outside the null distribution exceeds threshold (default 0.95). PlaceboInTime requires a PyMC model because it consumes posterior draws.
Walkthrough#
Fit the synthetic control as above so the experiment exposes a
treatment_timeandpost_impact.Run the pipeline cell below.
EstimateEffectre-fits the experiment inside the pipeline soPlaceboInTimecan clone the configuration for each placebo fold.Inspect the printed
CheckResult.textfor the verdict and per-fold summaries; the full hierarchical-null draws are available inmetadata["null_samples"].Open the HTML report (iframe) for a consolidated view alongside the main estimates.
Interpreting pass and fail. A pass means the actual cumulative impact is unlikely under the status-quo null inferred from placebo folds — consistent with a real treatment effect. A fail means the placebo folds produced effects of similar magnitude, so the real effect is hard to distinguish from background variability. Like any single diagnostic, this is necessary but not sufficient: passing does not prove identification, and failing does not prove the absence of an effect [Reichardt, 2019].
If this check fails#
A failing PlaceboInTime indicates that pseudo-interventions in the pre-period produced effects comparable to the real one. Concrete steps to diagnose and address this:
Inspect fold summaries. The
metadata["fold_results"]field on theCheckResultlists each placebo fold’s posterior mean and standard deviation. Look for a single noisy outlier versus a systematic pattern across folds — the latter is the stronger warning.Reconsider the donor pool. Highly correlated donors stabilise the synthetic control; donors with weak or negative correlation can introduce noise that shows up as spurious placebo effects. Re-run the pre-treatment correlation diagnostic above and prune the pool accordingly [Abadie, 2021].
Add complementary SC-specific checks. Run
LeaveOneOutto see whether any single donor drives the result, andPlaceboInSpaceto check whether spurious effects are common across control units. Persistent failures across multiple diagnostics are a stronger signal than any one alone.Sanity-check the convex-hull condition. Use
ConvexHullCheckto verify the treated unit’s pre-period trajectory lies within the support of the donor pool. Outside-the-hull fits depend on extrapolation and are more vulnerable to false positives [Abadie et al., 2010].Probe prior sensitivity. Re-fit with
cp.checks.PriorSensitivityusing tighter or more diffuse priors. If the conclusion flips, prior-data conflict is contributing to the fragile inference.
sensitivity_result = cp.Pipeline(
data=df,
steps=[
cp.EstimateEffect(
method=cp.SyntheticControl,
treatment_time=treatment_time,
control_units=["a", "b", "c", "d", "e", "f", "g"],
treated_units=["actual"],
model=cp.pymc_models.WeightedSumFitter(
sample_kwargs={"target_accept": 0.95, "random_seed": seed},
),
),
cp.SensitivityAnalysis(
checks=[cp.checks.PlaceboInTime(n_folds=2, random_seed=seed)],
),
cp.GenerateReport(include_plots=True),
],
).run()
for cr in sensitivity_result.sensitivity_results:
print(cr.text)
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [beta, y_hat_sigma]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 14 seconds.
Sampling: [beta, y_hat, y_hat_sigma]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [beta, y_hat_sigma]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 4 seconds.
Sampling: [beta, y_hat, y_hat_sigma]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
/Users/benjamv/git/CausalPy/causalpy/experiments/synthetic_control.py:118: UserWarning: Control units ['g' (r=-0.544)] have pre-treatment correlation below 0.0 or undefined with treated unit 'actual'. Consider excluding them from the donor pool. Use cp.plot_correlations() to inspect. See Abadie (2021) for guidance on donor pool selection.
self._check_donor_correlations()
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [beta, y_hat_sigma]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 10 seconds.
Sampling: [beta, y_hat, y_hat_sigma]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Sampling: [y_hat]
Initializing NUTS using jitter+adapt_diag...
Multiprocess sampling (4 chains in 4 jobs)
NUTS: [mu_status_quo, tau_status_quo, fold_z]
Sampling 4 chains for 1_000 tune and 1_000 draw iterations (4_000 + 4_000 draws total) took 1 seconds.
Sampling: [theta_new]
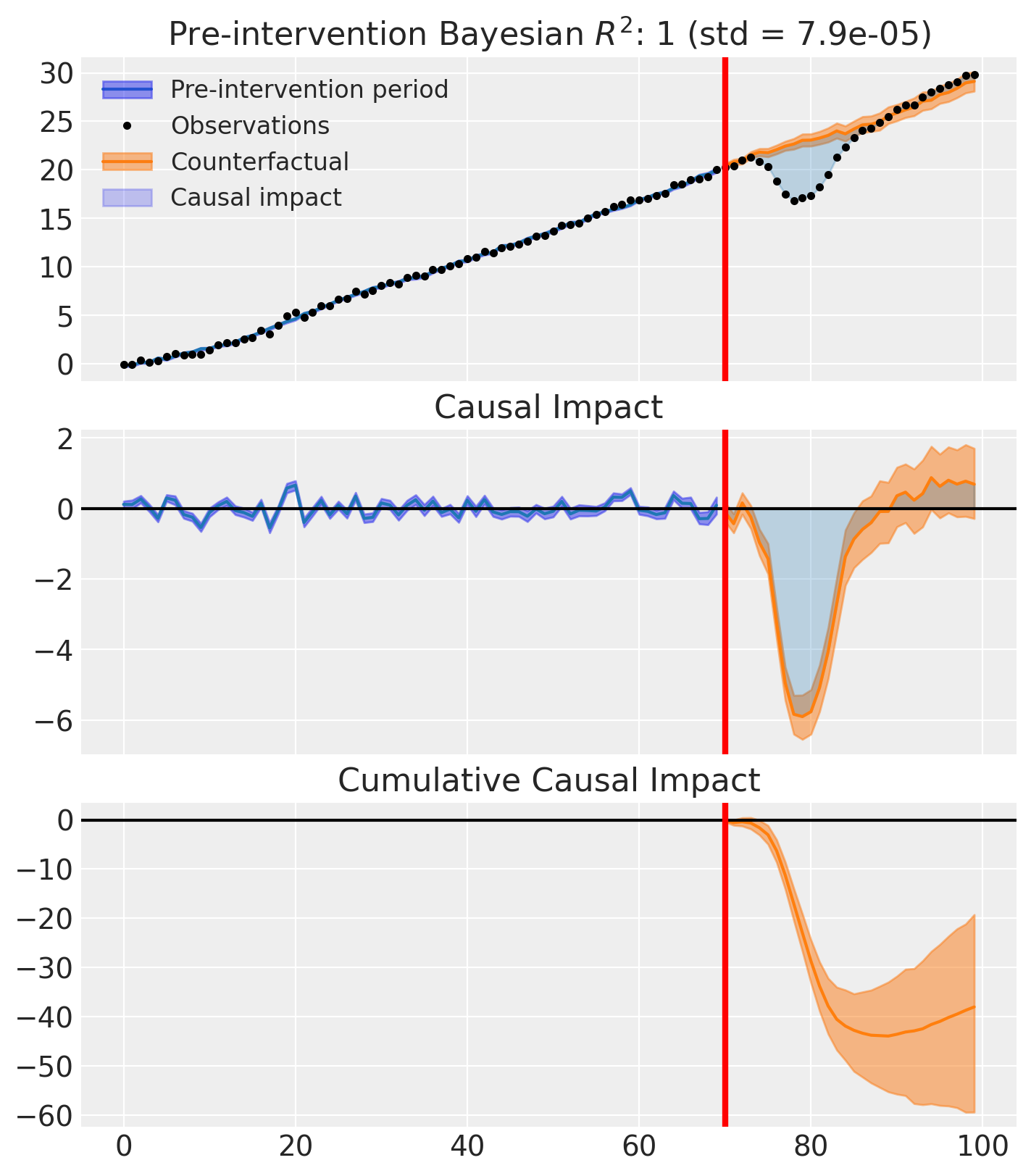
Placebo-in-time analysis: 2 of 2 folds completed.
Hierarchical status-quo model: mu=2.40, tau=12.48.
Actual cumulative impact: -38.00. P(actual outside null) = 0.918.
NOT SUPPORTED — actual effect is within the null distribution.
Fold 1: pseudo treatment at 12 — mean=16.71, sd=30.53
Fold 2: pseudo treatment at 41 — mean=-1.81, sd=9.18
import html as html_module
import warnings
from IPython.display import HTML
with warnings.catch_warnings():
warnings.filterwarnings(
"ignore", "Consider using IPython.display.IFrame", UserWarning
)
display(
HTML(
f'<iframe srcdoc="{html_module.escape(sensitivity_result.report)}" '
f'width="100%" height="600" style="border:1px solid #ccc;"></iframe>'
)
)

